The last months (years? since ever???) have seen a surge in populism and a rise in nationalism. Not only in Russia, the United States, Turkey, but also in some EU countries the ghost of nationalism-populism seems to be marching and gaining ground.
As to Germany, in September 24, 2017, the 19. German federal elections took place. The newly founded alt-right AfD (Alternative for Deutschland) has made a leap and moved in the Bundestag. In some electoral districts, its share rose to 35%, being the strongest party (although normally its share was lower), and in total, the AfD collect 12.6% of the votes, according to the official results.
Given the “hell and back” Europe and the world had to witness in the 20th century, this backlash seems surprising and alarming. Of course, a number of ideas on the origin of populist thought and countermeasures are out there.
What’s populism?
Populism is hard to define, see Laclau, On Populist Reason. So maybe authoritarianism and nationalism are concepts more adequate to deal with enemies of open society. Another idea would be to follow Popper’s idea of what makes a society open and what are the components of a “closed” society.
Without worrying much on ongoing discussions, I have tried to define and measure populism as manifest in Tweets of German politicians. The theoretical concept is not too much elaborated (as yet), I confess, but it builds on a well-known and intriguing source, Poppers Open Society.
Poppers idea of populism
Basically, the following theoretical aspects were used for gauging populism.
The roots of the (Western) civilization may be traced to societies such as the ancient Greek clans. In the beginning (but in parts, later on too), those clans were patriarchal in the sense that there one men or one family or one cast who was ruling without any democratic basis. Basically, someone was telling what to do. And every one had to follow. Even more, know only what to do but rather what was right and what was wrong. There is, as a consequence, no place for questions and questioning or doubt and doubting in such societies. It’s pretty clear and easy what’s right and wrong, and what you ought to now. Now, modern society is quite complex, and as a consequence, it is difficult to say what will happen, and what you should do. There are much less guides and less guidance as to what the “right” way of living consists of. Poppers argues that people are suffering from this insecurity, albeit to a different degree. Especially in times of crisis, rapid change, and strong progress the desire too come back to “good old days”, with all their simplicity and security offered by strong men and strong ideas (in the sense of that they provide sure answers), rises.
Moreover, those old tribal communities consisted (often) of kinship, there were genetic similarity among the members of a tribe, compared to the people of other tribes. As a result, such tribes or rather such tribal thinking rejoices in evoking the “blood is thicker than water” ideas. You should prefer your relatives more than strangers; sounds plausible, doesn’t it? But taken farer this leads a walling-off from the outside, in the sense that outsiders are considered negative or eve as enemies. In short, the “we” as a group - family, caste, tribe, culture, people, folk - should hold together, and should regard outsiders with suspicion.
Similarly, not only our “blood” but also our “soil” is what defined (defines) us as tribal society. This is our land. Not yours. We have always, okay, at least for a long time, been inhabiting this land.
More coherently, this is our group, our land, our stories. Nobody (including myself and you, as possible parts of this group) may object to this notion. We may not even doubt, we must obey to the law of what the strong man says, and he says he knows what to know, that’s why he is strong. Admitting “I don’t know” would be considered as a sign of weakness, not of strength. Don’t propose that things could be like this, but also like that, and even this may vary from time to time or according to some unknown boundary conditions…
Such ideas provide the base of “closed society thinking”, and populism is nothing more than alluding to this thinking and supporting it.
Indicators of populism in the present research
Much more down to earth, and certainly cum grano salis to say the least, I have tried to translate Popper’s ideas into these eight indicators:
- word length or rather word brevity
- Number of negative connotated words vs. number of positively connotated words (odds)
- Proportion of negatively connotated words
- Proportion of emotional words
- Score (intensity) of emotion
- Score (intensity) of negative emotion
- Proportion of words in CAPITAL LETTERS (shouted)
- Number of adjectives vs. number of adverbs
Out of these indicators, I formed one populism measure, and I then compared political parties based on this measure. I did not want to tap too much into the individual persons, partly because an aggregated measure may be more reliable, and partly because not too be to nasty to individual persons.
Data material
I collected around 400k tweets via the Twitter API; plus approx. 30k Tweets by Donald Trump. In sum, about 200 politicians donated about 6 millions words for this analysis. Not all parties are included, only seven important, the remainder is ignored (sorry). Data was collected from several years, as provided by the API.

I think that the API prefers providing newer tweets. Note that Trump data has been accessed from the Trump Twitter Archive.
Not all parties tweet the same amount. As I have recently read on Twitter, the only 12 hours, Donald Trump did not tweet was the day he was elected President…
From some parties, I was able to find a lot of accounts, from other parties, not so many. This may of course provide some basis for bias. I have tried to find some overview or “official” list of politicians’ twitter accounts. What I found was this. In addition, I added the accounts from the Bundesvorstaende of the AfD and the FDP, because these parties were especially lacking accounts.

Reproducibility
The code is on Github, completely. In this post, I will not discuss all technical aspects, but rather invite everyone interested to read the code.
Tweets per day
Who is the greatest… tweep? Who “does it” most frequently? Well, not too much surprising, Donald Trump is ahead of the pack (again).
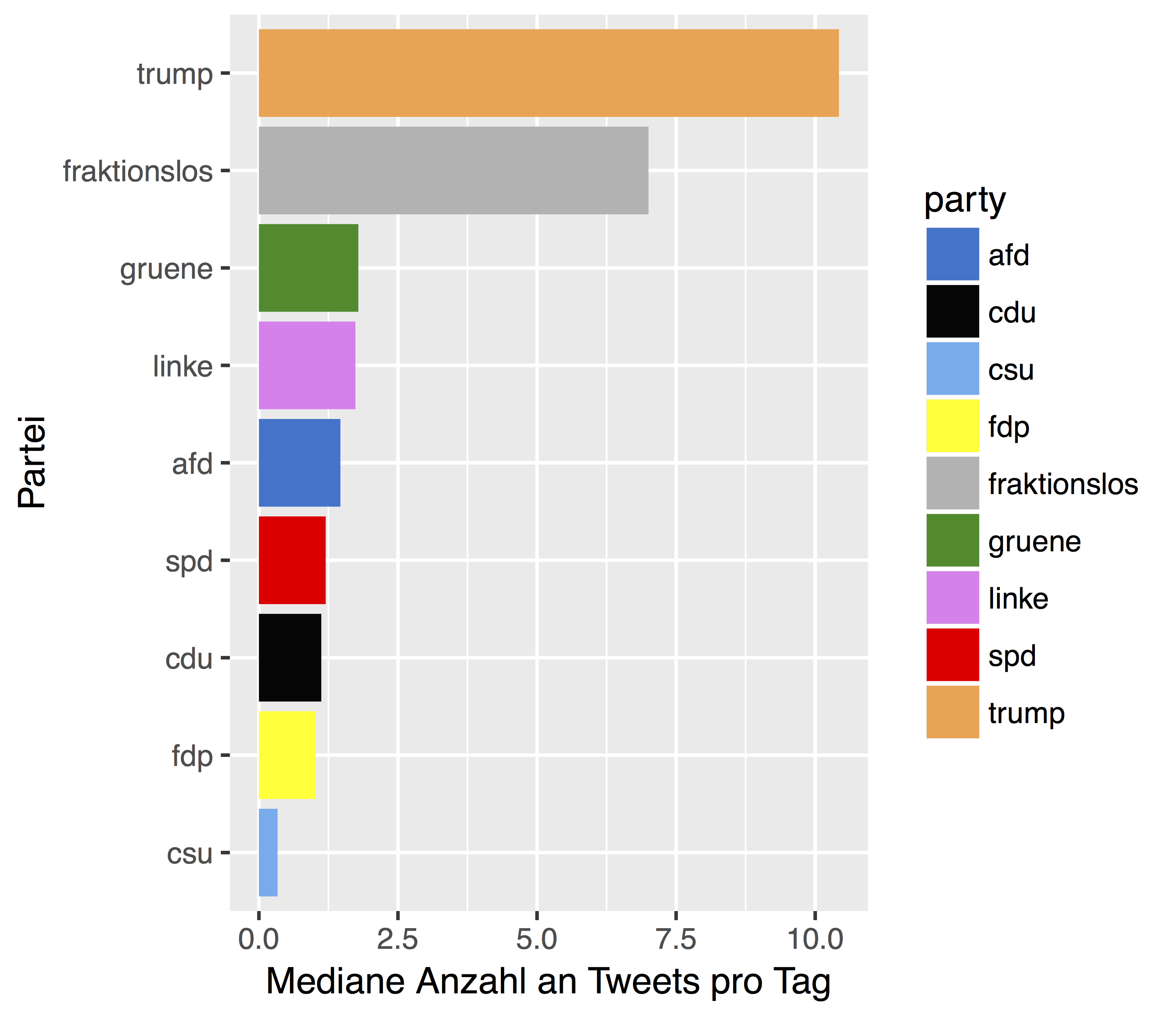
But considering only the German parties, we find that the Greens are leading.

Populism score
Averaged (by median) over the 8 indicators, one finds that the populism scores are … not too much of a surprise, in some regard.

Trump gets the largest part, the highest score. But maybe it would be better to leave him out of this “ranking”, as the different languages (German vs. English) cannot readily be compared with regard to such nuanced things like populism. Anyhow, the two most extreme parties, the AfD (alt-right) and the Linke (Leftists) are the ones with highest populism scores. Not that zero has been centered as the median over all accounts.
Cave and conclusion
Whereas Poppers theory certainly is compelling, the choice of indicators remains subjective. This is not unique for the present analysis, but still different sets of indicators may provide different pictures. Still, this picture appears well backup by the data. What’s your impression, your thoughts? Feel free to discuss your ideas.